Anti-CRISPR Proteins: A Biological Safety Switch for Precise Gene Editing
Fatemeh Akhoondi, MSc, Molecular and Cellular Biology, Independent Researcher
CRISPR has become a powerful tool for gene editing, yet achieving safe and precise control remains challenging. Anti-CRISPR (Acr) proteins provide a natural way to slow or stop Cas enzymes when needed. By reducing off-target activity, maintaining a short editing window, enabling tissue-specific control, and enhancing delivery consistency, these proteins provide an additional layer of regulation. This article examines the mechanisms by which these proteins function and discusses their significance for developing safer, more precisely controlled gene-editing systems.
Introduction
The Missing Safety Layer in Gene Editing
The rapid growth of CRISPR-based gene editing has created significant opportunities in medicine, biotechnology, and therapeutic development. These technologies are now essential for next-generation therapies, including the treatment of monogenic disorders and the engineering of immune cells.
However, they also pose safety risks, including off-target edits, prolonged nuclease activity, immune responses, and uncontrolled editing windows. Transitioning CRISPR technology from experimental applications to widespread clinical use requires the development of robust and programmable safety mechanisms. Anti-CRISPR proteins (Acrs), which are natural inhibitors of CRISPR systems, represent a particularly promising solution. Anti-CRISPR proteins act as biological “off switches” that enable precise, temporal regulation of CRISPR activity. Initially identified in bacteriophages as mechanisms to evade bacterial immune defenses, these proteins are currently being adapted to address significant challenges in therapeutic gene editing (1,2).
Why Enhanced Safety Control Matters in CRISPR Therapies
CRISPR-based therapeutics have entered clinical trials at a rapid pace, especially in fields such as:
- AAV gene therapy
- CAR-T and engineered immune cells
- Ex vivo CRISPR editing of hematopoietic cells
- In vivo base editing and prime editing
- Somatic gene-correction therapies
However, these technologies face well-documented obstacles:
- Off-target edits: Off-target effects arise when CRISPR cleaves DNA sequences that partially match the guide RNA, resulting in unintended genetic modifications.
- Persistent nuclease activity: If Cas9 remains in the cell longer than needed, it may cause additional, unpredictable edits and increase genotoxicity.
- Immune activation: Cas9 derived from bacteria such as Streptococcus pyogenes may trigger adaptive immune responses.
- Lack of precise temporal control: Traditional modulation methods, such as dose variation, transient delivery, and chemical switches, do not offer precise, reliable timing of CRISPR activity.
A safe therapeutic platform requires precise biological control of CRISPR activity. Anti-CRISPR proteins offer this unique capability.
How Anti-CRISPR Proteins Work
Anti-CRISPR proteins are small, phage-derived inhibitors that interfere with CRISPR-Cas systems through several distinct molecular mechanisms (3):
- Blocking DNA binding: These proteins prevent Cas enzymes from interacting with their target DNA.
- Inhibiting nuclease activity: In this mechanism, Cas enzymes bind to DNA but are unable to cleave it, resulting in a non-functional or “silent” nuclease.
- Disrupting sgRNA loading: This mechanism prevents the assembly of functional CRISPR-Cas complexes.
- PAM mimicry: This strategy involves decoying Cas enzymes to bind the inhibitor rather than the genomic DNA.
Different anti-CRISPR proteins target distinct structural domains, offering a diverse molecular toolkit for programmable inhibition of genome editing. Recent reviews indicate that more than 100 validated Acr proteins have been identified across multiple CRISPR types, each exhibiting unique inhibitory mechanisms (1). This natural diversity provides a modular set of inhibitors that researchers can tailor to therapeutic needs.
Anti-CRISPR as a Safety Switch in Cell & Gene Therapies
1. Enhancing CAR-T and immune cell engineering
CAR-T therapies require precise editing to prevent oncogenic rearrangements, abnormal receptor expression, and chromosomal fusions. Anti-CRISPR proteins introduce programmable time limits, functioning as temporary brakes that reduce risk without affecting therapeutic potency. Studies in mammalian systems show that delayed Acr delivery (e.g., AcrIIA4 delivered 6 hours post-Cas9) significantly enhances editing precision and reduces off-target mutagenesis without affecting therapeutic viability (1).
2. Improving the Safety of Ex Vivo Stem Cell Editing
Ex vivo Hematopoietic stem cells (HSCs), editing is highly sensitive to even small perturbations in genomic integrity. Anti-CRISPR proteins can address these challenges in several ways:
- They help tighten the editing window,
- Improve on/off-target discrimination
- Reduce prolonged editing exposure
- Enhance the safety profile of engrafted cells
3. Increasing Control in AAV-Based Gene Therapies
AAV-delivered CRISPR remains active for extended periods, potentially increasing the risk of off-target effects. Anti-CRISPR proteins can be introduced via co-expression, independent vectors, or synthetic gene circuits to restrict this duration.
Tissue-specific control is possible with miRNA-responsive Acr constructs. For example, AcrIIC3 regulated by miR-122 enables liver-specific Cas9 inhibition in mice (4).
4. Programmable On/Off for Base Editors & Prime Editors
Base editors and prime editors act for extended durations, raising risks of unintended deamination. Anti-CRISPRs can address these risks in several ways:
- Pausing or terminating the editing process when needed
- Enabling tighter control over exposure duration
- Improving overall editing precision
AcrIIA2, AcrIIA4, and AcrIIA5 have been shown to inhibit both adenine and cytosine base editor systems in a dose-dependent manner, reducing off-target editing in human cells (5) (6).
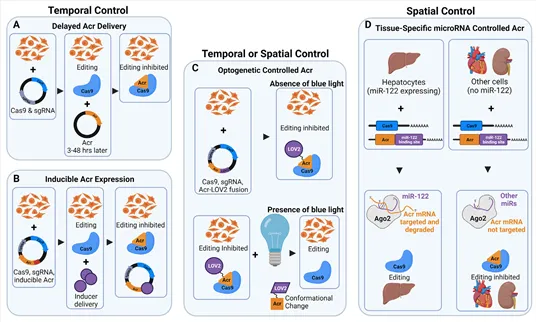
Figure 1 Overview of anti-CRISPR strategies for temporal and spatial control of genome editing.
Several approaches can modulate when and where CRISPR–Cas9 is active: (A) Delaying Acr delivery allows editing to occur first, followed by timed inhibition. (B) Acr expression can be switched on later using inducible promoters or chemical triggers. (C) Fusions with light-responsive domains enable optogenetic control, where illumination determines Acr activity in specific regions or time windows. (D) Tissue-restricted microRNAs can selectively suppress Acr transcripts, providing organ- or cell-type–specific regulation of Cas9 (1).
Engineering Synthetic Anti-CRISPR Circuits
Advances in synthetic biology enable the integration of Acrs into engineered biological circuits, including:
- Drug-inducible switches
- Optogenetic (light-controlled) Acrs
- MicroRNA-specific Acr control
- AAV-compatible regulatory modules
Optogenetic systems such as CASANOVA (AcrIIA4-LOV2 fusion) enable reversible, light-activated CRISPR inhibition. This approach offers one of the most precise control strategies developed to date.
MicroRNA-driven Acr repression (e.g., miR-122 restricted to hepatocytes) allows spatial control by coupling Acr expression to tissue-specific microRNA signatures (7) (1).
Future Outlook
Although anti-CRISPR proteins are relatively new to therapeutic development, their potential is becoming clearer as genome-editing technologies advance. As CRISPR systems are applied in more complex clinical settings, the need for reliable mechanisms to fine-tune or temporarily halt editing activity is expected to increase. Anti-CRISPR proteins offer a natural basis for this control, and several promising areas of progress are emerging.
One ongoing direction involves the development of engineered or optimised Acr variants with improved stability and reduced immunogenicity for use in human cells. These efforts build on the structural understanding of how natural Acrs interact with different Cas enzymes and represent a gradual, evidence-based extension of current research. Scientists are also investigating ways to connect Acr activity to regulatory elements, such as protein domains responsive to small molecules, cellular stress, or physiological conditions. Although these strategies are in early development, they offer practical options to enhance the precision and control of genome-editing tools.
Advances in delivery technologies will affect how efficiently Acrs are incorporated into therapeutic programs. Enhanced protein delivery, improved viral and non-viral vectors, and targeted nanoparticle platforms may increase the effectiveness of co-delivering or sequentially delivering Acrs with CRISPR components in vivo. As these systems develop, integrating anti-CRISPR proteins into experimental and clinical workflows will likely become more practical.
Overall, these steady advancements indicate that anti-CRISPR proteins will remain key components of precision gene-editing systems, especially where controlled, time-limited, or reversible editing is needed for safety and therapeutic reliability.
Conclusion
Anti-CRISPR proteins constitute a significant emerging layer of regulation for CRISPR-based therapeutics. Through reversible, adjustable, and context-specific inhibition of nuclease activity, these proteins address critical safety concerns related to genome editing. Advances in elucidating Acr mechanisms, enhancing protein engineering, and optimising delivery strategies are steadily increasing their practical utility. As clinical applications of CRISPR expand, anti-CRISPR systems are anticipated to play a progressively supportive role in the development of genome-editing approaches that are safer, more predictable, and better suited to therapeutic needs.
References
- Kraus C, Sontheimer EJ. Applications of Anti-CRISPR Proteins in Genome Editing and Biotechnology. Vol. 435, Journal of Molecular Biology. Academic Press; 2023.
- Davidson AR, Lu WT, Stanley SY, Wang J, Mejdani M, Trost CN, et al. BI89CH13_Davidson ARjats.cls Anti-CRISPRs: Protein Inhibitors of CRISPR-Cas Systems. 2025;26:46. Available from: https://doi.org/10.1146/annurev-biochem-011420-
- Li Y, Bondy-Denomy J. Anti-CRISPRs go viral: The infection biology of CRISPR-Cas inhibitors. Vol. 29, Cell Host and Microbe. Cell Press; 2021. p. 704–14.
- Lee J, Mou H, Ibraheim R, Liang SQ, Liu P, Xue W, et al. Tissue-restricted genome editing in vivo specified by microRNA-repressible anti-CRISPR proteins. 2019; Available from: http://www.rnajournal.org/cgi/doi/10.1261/rna.
- Zhao M, Gao A, Zhu Y. Research Progress of Phage Anti-CRISPR Proteins.
- Marino ND, Pinilla-Redondo R, Csörgő B, Bondy-Denomy J. Anti-CRISPR protein applications: natural brakes for CRISPR-Cas technologies. Vol. 17, Nature Methods. Nature Research; 2020. p. 471–9.
- Zhang Y, Marchisio MA. Type II anti-CRISPR proteins as a new tool for synthetic biology. Vol. 18, RNA Biology. Bellwether Publishing, Ltd.; 2021. p. 1085–98.










